antibodies
▼: adhesion molecules : antigen binding site : C : CH1-4 : cellular adhesion molecules : complement fixation : complementarity determining regions : constant domains : domains : evolution of immunoglobulins : Fab : Fc : heavy chain : hinge region : Ig supergene family : isotypes : kinase activation : light chain : location of Ig classes : membrane-bound Igs : multimeric structures : tissue location : V : VDJ recombination : VH, VL : variable domains :▼
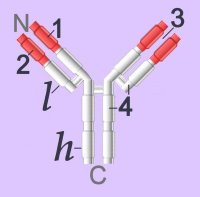 Immunoglobulins (left - click to enlarge) comprise two heavy (h) and two light-chain (l) protein subunits, each of which folds into domains (4 on heavy, 2 on light). These adhesion sites or domains contain one or more folds of 60 to 100 amino acids.
Immunoglobulins (left - click to enlarge) comprise two heavy (h) and two light-chain (l) protein subunits, each of which folds into domains (4 on heavy, 2 on light). These adhesion sites or domains contain one or more folds of 60 to 100 amino acids.Depending upon the character of the heavy chain, immunoglobulins are divided into five classes – IgG, IgD, IgE, IgA, IgM – that are expressed in different tissues. The classes are further subdivided into isotypes, which have different properties in terms of complement fixation and binding to immunoglobulin (Ig) receptors.
Members of the immunoglobulin supergene family are found as:
● membrane-bound surface receptors of immune-system cells,
● cellular adhesion molecules, or
● soluble antibodies (γ-globulins) synthesized by activated B cells.
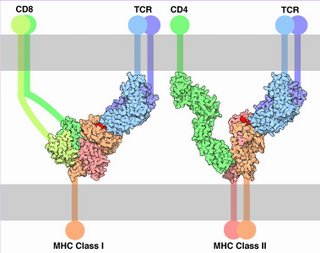
Membrane-bound Igs have a transmembrane segment and a cytoplasmic C-terminal tail. The 2 β- chains are stabilized into sandwiched β sheets that are adherent by virtue of hydrophobic interactions between disulphide bonds. Igs assume a Y-shaped structure "topped" at the extracellular N-terminals by variable domains (red), with a variable domain at the tip of the heavy chain (1) and the light chain (2), between which lies an antigen binding site (3). The variable regions are coded by pluripotential DNA sequences that can generate thousands of polypeptide sequences capable of adhering to millions of different ligands. Binding is homophilic or heterophilic, including binding to different Igs and to integrins. Both light and heavy chains contain constant domains (white, 4).
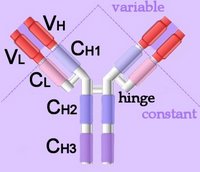 Right - click to enlarge - the heavy chains of IgA, IgD and IgG each have four domains, where those at the N-terminal are variable (VH) and the other three are constant (CH1-3). IgE and IgM have one variable and four constant domains (CH1-4) on the heavy chain. The variable domains are termed Fab, while the constant domains are termed Fc.
Right - click to enlarge - the heavy chains of IgA, IgD and IgG each have four domains, where those at the N-terminal are variable (VH) and the other three are constant (CH1-3). IgE and IgM have one variable and four constant domains (CH1-4) on the heavy chain. The variable domains are termed Fab, while the constant domains are termed Fc.The light chains have two domains, one variable domain (VL) at the N-terminal, and one constant (CL) domain.
The antigen binding site lies between VH and VL (shaded lavendar). Most variability is found in three superficial-loop forming regions in the VH and VL domains, which are the complementarity determining regions or CDRs. CDR3 binds antigens and CDR1-2 bind MHCs. CDR3 shows more variation that do either CDR1 or 2.
The domains have related amino acid sequences that possess a common secondary and tertiary structure. This conserved structure is found frequently in proteins involved in cell-cell interactions and is particularly important in immunology. The constant (Fc) regions have complement fixing and Ig receptor binding activity. The hinge region, in IgG, IgA and IgD, is an important sequence of 10-60 amino acids between CH1 and CH2 that confers flexibility on the molecule.
animations Џ B cell selection Џ ELISA test +ve, -ve Џ IgG rotating x- y- axes Џ Rotating mouse IgG2a Molecule (y-axis) Џ somatic recombination of Ig gene Џ spinning IgG1 Kol Џ unfolding (small) IgG . unfolding (large) IgG .
Immunoglobulins attain their enormous variability by splicing components (VDJ recombination) coded in widely scattered sequences of DNA that are located in two different chromosomes. Antigen binding takes place at the heavy chain, which displays enormous variation by virtue of combining 1 of 400 possible variable gene segments with 1 out of 15 diversity segments and 1 out of 4 joining segments. This alternative splicing generates 24,000 possible combinations for the DNA encoding the heavy chain alone. The variable coding segments are assembled together with those for the constant-C segments of the heavy-chain molecule.
Tissue location:
IgA – mucus – gut, respiratory tract
IgD – antigen receptor on B cells
IgE – mast cells – releases histamines in response to allergens
IgG – primary immunity against invading pathogens
IgM – early B cell-mediated response to invading pathogens
Some antibody classes form multimeric structures – pentamers (IgM) and dimers or trimers (IgA). These two isotypes also associate with a small protein called the joining (J) chain required for stabilisation of the complexes.
The immunoglobulin superfamily is evolutionarily ancient, is widely expressed, and is constitutive or long-term up-regulated. Immunoglobulin antibodies are released by activated B cells of the immune system, on which they also act as surface marker proteins. Adherence of immunoglobilins to foreign substances or to cellular invaders may be sufficient to disarm the invader, or the attached antibodies function as attack signal to macrophages and natural killer cells. Adhesion molecules of the immunoglobulin supergene family, activate specific kinases through phosphorylation, resulting in activation of transcription factors, increased cytokine production, increased cell membrane protein expression, production of reactive oxygen species, and cell proliferation.
▲: adhesion molecules ~ adhesion molecules ф antigen : antigen binding site ф APCs ф B cells : C : CH1-4 : cellular adhesion molecules : complement fixation ф complement system : complementarity determining regions : constant domains : domains : evolution of immunoglobulins : Fab : Fc : heavy chain : hinge region ф humoral immunity : Ig supergene family ~ immunoglobulins : isotypes : kinase activation ♦ kinases : light chain : location of Ig classes : membrane-bound Igs : multimeric structures ф receptors ф signaling ф surface receptors ф T cells : tissue location ~ tyrosine kinases : V : VDJ recombination ф VDJ recombination : VH, VL : variable domains :▲
Tables Fc receptors Immune Cytokines Immunoglobulins Cell Adhesion Molecules Cell signaling Receptor Tyrosine Kinases (RTKs) Receptor Signal Transduction Second Messengers
▲ Top ▲
tags [Immunology]
[antibodies]
Labels: adhesion molecules, antibodies, antigen binding, complementarity determining regions, domains, evolution, heavy chain, immunoglobulins, isotypes, surface marker proteins, VDJ recombination
| 0 Guide-Glossary